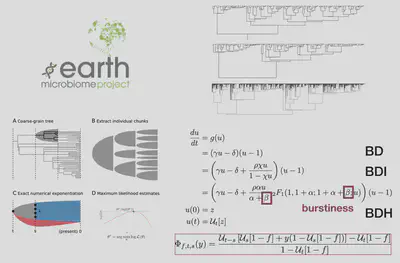
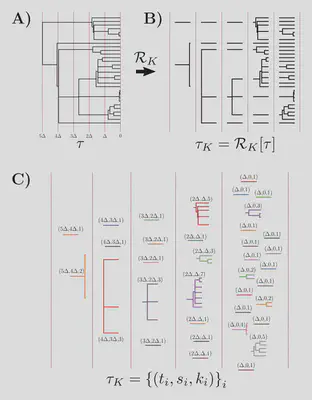
Microbial diversification dynamics
I developed a coarse-grained inference framework to infer the tempo, mode, and shape of the global microbial tree of life using a novel diversification model called the birth, death, and heterogeneous innovation model. This approach has three favorable features. First, the model makes an explicit effort to acknowledge and capture empirical features of microbial phylogenies such as their “burstiness” and the effective lengthening of ancestral lineages due to unobserved extinctions and incomplete sampling. Second, the introduction of the heterogeneous innovation process overcomes the problem of parameter non-identifiability plaguing more traditional speciation-extinction models between the parameters characterizing the sampling fraction and the speciation rate. Third, the coarse-graining step overcomes the unavoidable problem of incompletely resolved phylogenies common to microbial datasets. Using this approach, I identified a previously unknown universality pattern inherent to microbial diversification. This universality manifests itself across a vast range of different microbiome through the appearance of a common exponent, ranging between 1.3 to 1.7, characterizing the tail of the burst size distribution in their phylogenies.