Inference of tree-structured microbial interactions
In this project we explore the possibility of inferring full interaction matrices within a community (in our case microbial) using only O(1) community samples. This dramatic reduction in the size of the required dataset is made possible using ecologically and biologically justified assumptions. On one hand we assume that the shape of the interaction matrix is tree-structured, namely that it aligns itself with the underlying phylogeny between the species of the community. On the other hand we assume that the community is forever embedded in a stochastic environment which constantly perturbs it at random intervals. This latter assumption thus that the community is never observed at equilibrium.
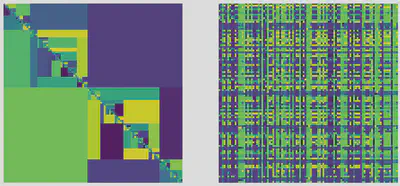
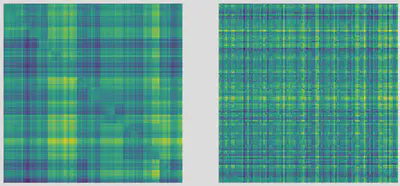
The two figures below are visual representation of random asymmetric tree-structured interaction matrices following two different scenario for the way interactions have evolved through time. The first one assumes interaction strengths are dominated by the original innovation or nature of the geological event that led to a speciation event. The second one assumes that interaction strengths are sums of accumulations of evolutionary differences along the paths in the phylogeny.